Deconvolution in Optical Microscopy
Deconvolution is a computationally intensive image processing technique that is being increasingly utilized for improving the contrast and resolution of digital images captured in the microscope. The foundations are based upon a suite of methods that are designed to remove or reverse the blurring present in microscope images induced by the limited aperture of the objective.
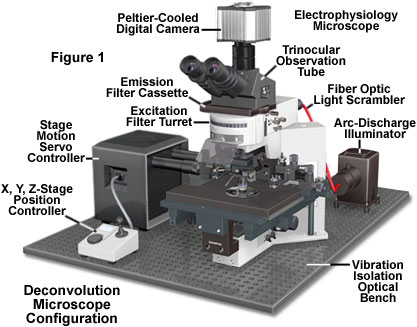
Practically any image acquired on a digital fluorescence microscope can be deconvolved, and several new applications are being developed that apply deconvolution techniques to transmitted light images collected under a variety of contrast enhancing strategies. One of the most suitable subjects for improvement by deconvolution are three-dimensional montages constructed from a series of optical sections.
Introduction to Deconvolution - As a technique, deconvolution is often suggested as a good alternative to the confocal microscope. This is not strictly true because images acquired using a pinhole aperture in a confocal microscope can also be analyzed by deconvolution techniques. However, a majority of the deconvolution experiments reported in the literature apply to images recorded on a standard widefield fluorescence microscope. Modern deconvolution algorithms have yielded images of comparable resolution to that of a confocal microscope. In fact, confocal microscopy and widefield deconvolution microscopy both work by removing image blur, but do so by opposite mechanisms.
Algorithms for Deconvolution Microscopy - Over the past ten years, a wide variety of both simple and complex algorithms have been developed to assist the microscopist in removing blur from digital images. The most commonly utilized algorithms for deconvolution in optical microscopy can be divided into two classes: deblurring and image restoration. Deblurring algorithms are fundamentally two-dimensional, because they apply an operation plane-by-plane to each two-dimensional plane of a three-dimensional image stack. In contrast, image restoration algorithms are properly termed "three-dimensional" because they operate simultaneously on every pixel in a three-dimensional image stack.
Artifacts and Aberrations in Deconvolution Analysis - After deconvolution algorithms have been applied, the restored image may include apparent artifacts such as striping, ringing, or discontinuous cytoskeletal staining. In some instances, these problems are related to data representation and will not occur with a different algorithm or software package. The artifacts can also occur when processing parameters are not correctly configured for the raw image. Finally, artifacts are often not caused by computation, but by histology, optical misalignment, or electronic noise. When attempting to diagnose the source of an artifact, the first step is to carefully compare the raw image with the deconvolved image.
Resolution Criteria for Deconvolution Analysis - Resolution in optical microscopy is often assessed by means of an optical unit termed the Rayleigh criterion, which was originally formulated for determining the resolution of two-dimensional telescope images, but has since spread into many other arenas in optics. The Rayleigh criterion is defined in terms of the minimum resolvable distance between two point sources of light generated from a specimen and is not dependent upon the magnification used to produce the image. In deconvolution analysis, the three-dimensional nature of the point spread function must be considered when applying the Rayleigh criterion.
Selected Literature References and Internet Resources
Deconvolution Resources on the Internet - Deconvolution is emerging as a powerful tool for high-resolution three-dimensional analysis of complex biological specimens. Assisting in the development of existing and new deconvolution techniques is a host of universities, research institutions, and commercial organizations. This section contains links to web resources on deconvolution analysis, including software packages, hardware (microscopes and accessories), and laboratories involved with the technology.
ZEISS Campus Deconvolution Microscopy Reference Library - Over the past several decades, deconvolution microscopy has become a mainstream image processing tool for deciphering the substructure of living and fixed specimens in three dimensions. Routinely applied to widefield optical sections, as well as those obtained in confocal and structured illumination, the technique has benefited from the continued development of advanced algorithms and turnkey systems. The references listed in this section point to review articles that should provide the starting point for a thorough understanding of deconvolution.
Literature Reference Listings - A number of review articles on deconvolution in optical microscopy have been published by leading researchers in the field and were utilized as references to prepare discussions included in this Digital Image Processing Section of the Molecular Expressions Microscopy Primer. This section contains periodical location information about these articles, as well as providing a listing of selected original research reports and books describing deconvolution as it pertains to digital image processing in optical microscopy.
Contributing Authors
Wes Wallace - Department of Neuroscience, Brown University, Providence, Rhode Island 02912.
Lutz H. Schaefer - Advanced Imaging Methodology Consultation, Kitchener, Ontario, Canada.
Jason R. Swedlow - Division of Gene Regulation and Expression, School of Life Sciences Research, University of Dundee, Dundee, DD1 EH5 Scotland.
Thomas J. Fellers and Michael W. Davidson - National High Magnetic Field Laboratory, 1800 East Paul Dirac Dr., The Florida State University, Tallahassee, Florida, 32310.
BACK TO DIGITAL IMAGING IN OPTICAL MICROSCOPY
